THIS ARTICLE IS MORE THAN FIVE YEARS OLD
This article is more than five years old. Autism research — and science in general — is constantly evolving, so older articles may contain information or theories that have been reevaluated since their original publication date.
A novel method allows researchers to rapidly generate multiple stem-cell colonies with mutations linked to autism or other conditions1.
The method combines the gene-editing tool CRISPR with the ability to transform human stem cells into mature cells. It scales up the gene-editing process and uses custom software to check its efficiency.
Researchers have used CRISPR to deactivate genes in human stem cells: The tool cuts DNA strands, and the cell repairs the break by adding or deleting DNA letters. But researchers can’t predict which repair process a cell will use. They then coax the edited stem cells into becoming mature cells, such as neurons.
However, this method cannot efficiently target more than one gene in a single experiment, so studying the effects of different genetic variants is time-consuming. One hurdle is verifying whether the cell creates these mutations by deleting or inserting DNA. Deletions often have different effects on cells than insertions.
The new method, described 10 October in Stem Cell Reports, allows researchers to edit different genes in many stem cells at the same time.
Scaling up:
The researchers first generated guide RNAs to direct the CRISPR enzymes to specific locations in the DNA. They delivered the RNAs and enzymes into human stem cells along with a green fluorescent protein. Cells that incorporate CRISPR edits emit a green glow.
The researchers isolated the glowing cells and placed a single cell into each well of a 96-well plate. The cells in each well form colonies of what should be genetically identical ‘clones.’
The researchers extracted and sequenced DNA from each colony and used dedicated software to analyze the sequencing data. The software calculates the proportion of cloned cells that carry the desired edit and reveals whether the edit is a deletion or an insertion.
The team applied the method to 5,000 colonies from two stem cell lines derived from different human embryos. For one cell line, about 40 percent of clones had the desired mutation. For the other, the proportion was 25 percent.
CRISPR introduced a DNA deletion 85 percent of the time and an insertion 15 percent of the time, the researchers found. Their software predicts how each mutation affects gene expression.
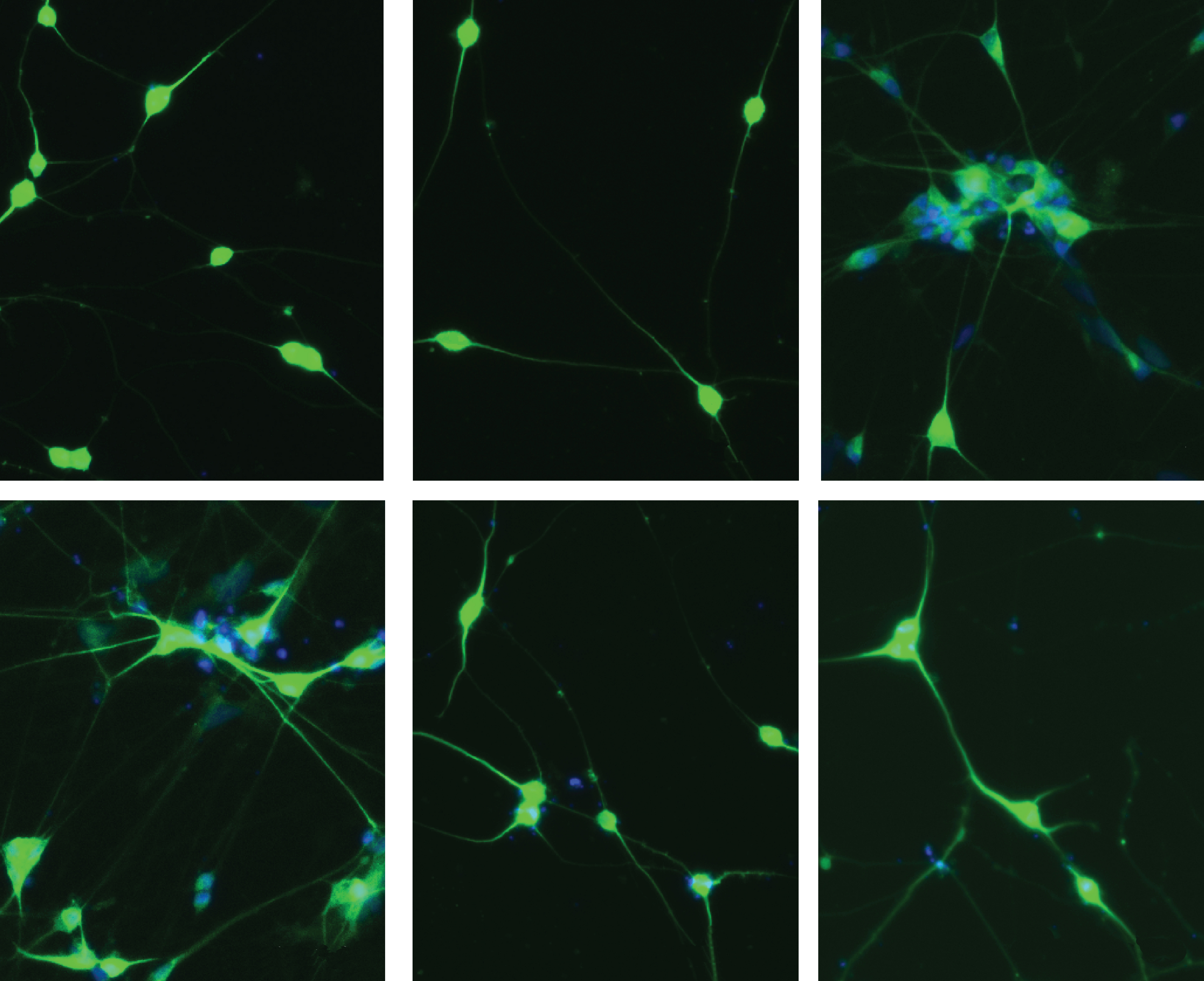
In one experiment, the researchers used the method to disrupt each of 58 genes related to schizophrenia, autism or intellectual disability in human stem cells. They then directed the cells to develop into neurons. The paper provides instructions for making the 58 guide RNAs used to target these genes.
By joining the discussion, you agree to our privacy policy.